Documentos de Académico
Documentos de Profesional
Documentos de Cultura
Transcripción 1
Cargado por
brxd69cmx7Derechos de autor
Formatos disponibles
Compartir este documento
Compartir o incrustar documentos
¿Le pareció útil este documento?
¿Este contenido es inapropiado?
Denunciar este documentoCopyright:
Formatos disponibles
Transcripción 1
Cargado por
brxd69cmx7Copyright:
Formatos disponibles
5¿ • Eukaryotic promoters are more varied than bacterial
promoters. Not only are different types of promoters
employed for the three polymerases, but there is great
variation within each type—especially among the ones
for protein-coding genes. Furthermore, some eukaryotic
RNA Hairpin loop
promoters are actually located downstream from the tran-
(GC-rich) scription start site.
• Binding of eukaryotic RNA polymerases to DNA requires
UUUUUU 3¿
the participation of additional proteins, called transcrip-
tion factors. Unlike the bacterial sigma factor, eukaryotic
transcription factors are not part of the RNA polymerase
A A A A
U UU U A A
molecule itself. Rather, some of them must bind to DNA
3¿ U 5¿ before RNA polymerase can bind to the promoter and
U
DNA initiate transcription. Thus, transcription factors, rather
5¿ 3¿
than RNA polymerase itself, determine the specificity
of transcription in eukaryotes. In this chapter, we limit
Self- our discussion to the class of factors that are essential
RNA
complementary
sequence
polymerase for the transcription of all genes transcribed by an RNA
polymerase. (We defer discussion of the regulatory class
of transcription factors, which selectively act on specific
genes, until Chapter 20.)
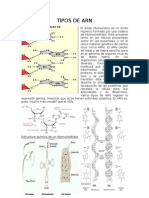
Figure 18-15 Termination of Transcription in Bacterial • Protein-protein interactions play a prominent role in the
Genes That Do Not Require the Rho Termination Factor. A first stage of eukaryotic transcription. Although some
short self-complementary sequence near the end of the gene transcription factors bind directly to DNA, many attach
allows the newly formed RNA molecule to form a hairpin loop struc- to other proteins—either to other transcription factors or
ture that helps dissociate the RNA from the DNA template. to RNA polymerase itself.
• RNA cleavage is more important than the site where tran-
scription is terminated in determining the location of the
In contrast, RNA molecules that do not form a GC-rich 3′ end of the RNA product.
hairpin loop require participation of the rho factor for termi-
• Newly forming eukaryotic RNA molecules typi-
nation. Genes encoding such RNAs were first discovered in
cally undergo extensive RNA processing (chemical
experiments in which purified DNA obtained from bacterio-
modification) both during and, to a larger extent, after
phage l was transcribed with purified RNA polymerase. Some
transcription.
genes were found to be transcribed into RNA molecules that
are longer than the RNAs produced in living cells, suggesting We will now examine these various aspects of eukaryotic
that transcription was not terminating properly. This problem transcription, starting with the existence of multiple forms of
could be corrected by adding rho factor, which binds to spe- RNA polymerase.
cific termination sequences 50–90 bases long located near the
3′ end of newly forming RNA molecules. The rho factor acts
as an ATP-dependent unwinding enzyme, moving along the RNA Polymerases I, II, and III Carry Out
newly forming RNA molecule toward its 3′ end and unwind- Transcription in the Eukaryotic Nucleus
ing it from the DNA template as it proceeds. Table 18-1 summarizes some properties of the three RNA
Whether termination depends on rho or on the formation polymerases that function in the nucleus of the eukaryotic
of a hairpin loop, it results in the release of the newly tran- cell, along with two other polymerases found in mitochondria
scribed RNA molecule and the core RNA polymerase. The core and chloroplasts. The nuclear enzymes are designated RNA
polymerase can then bind sigma factor again and reinitiate polymerases I, II, and III. As the table indicates, these enzymes
RNA synthesis at another promoter. differ in their location within the nucleus and in the kinds of
RNA they synthesize. The nuclear RNA polymerases also dif-
Transcription in Eukaryotic Cells Has fer in their sensitivity to various inhibitors, such as a-amanitin,
Additional Complexity Compared with a deadly toxin produced by the mushroom Amanita phalloides
Prokaryotes (the “death cap” fungus; the F-actin binding drug phalloidin,
introduced in Chapter 13, also comes from this organism).
Transcription in eukaryotic cells involves the same four stages
RNA polymerase I resides in the nucleolus and is re-
described in Figure 18-11, but the process in eukaryotes is
sponsible for synthesizing an RNA molecule that serves as a
more complicated than that in bacteria. The main differences
precursor for three of the four types of rRNA found in eukary-
are as follows:
otic ribosomes (28S rRNA, 18S rRNA, and 5.8S rRNA). This
• Three different RNA polymerases transcribe the nuclear enzyme is not sensitive to a-amanitin. Its association with the
DNA of eukaryotes. Each synthesizes one or more classes nucleolus is understandable because the nucleolus is the site
of RNA. of ribosomal RNA synthesis and ribosomal subunit assembly.
540
M18_HARD7694_09_GE_C18.indd 540 17/02/17 7:24 am
Table 18-1 Properties of Eukaryotic RNA Polymerases
RNA Polymerase Location Main Products A-Amanitin Sensitivity
I Nucleolus Precursor for 28S rRNA, 18S rRNA, and 5.8S rRNA Resistant
II Nucleoplasm Pre-mRNA, most snRNA, and microRNA Very sensitive
III Nucleoplasm Pre-tRNA, 5S rRNA, and other small RNAs Moderately sensitive*
Mitochondrial Mitochondrion Mitochondrial RNA Resistant
Chloroplast Chloroplast Chloroplast RNA Resistant
*In mammals.
RNA polymerase II is found in the nucleoplasm and
synthesizes precursors to mRNA, the class of RNA molecules Three Classes of Promoters Are Found in
that encode proteins. Rather than being diffusely distributed Eukaryotic Nuclear Genes, One for Each Type
throughout the nucleus, active molecules of polymerase II are of RNA Polymerase
located in discrete clusters, called transcription factories, that The promoters that eukaryotic RNA polymerases bind to are
may represent sites where active genes come together to be even more varied than bacterial promoters, but they can be
transcribed. In addition to producing mRNA precursors, RNA grouped into three main categories, one for each type of poly-
polymerase II synthesizes most of the snRNAs, small nuclear merase. Figure 18-16 shows examples of the three types of
RNAs involved in posttranscriptional RNA processing, and the promoters.
microRNAs, which regulate the translation and stability of The promoter used by RNA polymerase I—that is, the pro-
Chapter 18
specific mRNAs and, to a lesser extent, control the transcrip- moter of the transcription unit that produces the precursor for
tion of certain genes. Polymerase II is responsible for produc- the three largest rRNAs—has two parts (Figure 18-16a). The
ing the greatest variety of RNA molecules and is extremely part called the core promoter, defined as the smallest set of
sensitive to a-amanitin, which explains the toxicity of this DNA sequences able to direct the accurate initiation of tran-
|
compound to humans and other animals. scription by RNA polymerase, actually extends into the nucleo-
Gene Expression: I. The Genetic Code and Transcription
RNA polymerase II differs from polymerases I and III at its tide sequence to be transcribed. The core promoter is sufficient
C-terminus, where it has extra amino acids. The C-terminus for proper initiation of transcription, but transcription is made
of RNA polymerase II can be phosphorylated at a variety of more efficient by the presence of an upstream control element,
locations to produce what is sometimes called a phosphoryla- which for RNA polymerase I is a fairly long sequence similar
tion “code.” This “code” dramatically affects the functions of (though not identical) to the core promoter. Attachment of
polymerase II and correlates with where the enzyme is located transcription factors to both parts of the promoter facilitates
along the DNA as it continues transcription. As a result, this the binding of RNA polymerase I to the core promoter and en-
most versatile of the RNA polymerases is also the most tightly ables it to initiate transcription at the start site.
regulated. In the case of RNA polymerase II, at least four types of
RNA polymerase III is also a nucleoplasmic enzyme, DNA sequences are involved in core promoter function (Figure
but it synthesizes a variety of small RNAs, including tRNA 18-16b). These four elements are (1) a short initiator (Inr)
precursors and the smallest type of ribosomal RNA, 5S rRNA. sequence surrounding the transcriptional start site (which is
Mammalian RNA polymerase III is sensitive to a-amanitin often an A, as in bacteria); (2) the TATA box, which consists of
but only at higher levels of the toxin than are required to in- a consensus sequence of TATA followed by two or three more
hibit RNA polymerase II. (The comparable enzymes of some A’s, usually located about 25 nucleotides upstream from the
other eukaryotes, such as insects and yeasts, are insensitive start site; (3) the TFIIB recognition element (BRE) located
to a-amanitin.) slightly upstream of the TATA box; and (4) the downstream
Structurally, RNA polymerases I, II, and III are somewhat promoter element (DPE) located about 30 nucleotides
similar to each other as well as to bacterial core RNA poly- downstream from the start site. These four elements are or-
merase. The three enzymes are all quite large, with multiple ganized into two general types of core promoters: TATA-driven
polypeptide subunits and molecular weights around 500,000. promoters, which contain an Inr sequence and a TATA box
RNA polymerase II, for example, has more than ten subunits with or without an associated BRE, and DPE-driven promoters,
of at least eight different types. The three biggest subunits are which contain DPE and Inr sequences but no TATA box or BRE.
evolutionarily related to the bacterial RNA polymerase sub- Besides being found in eukaryotes, TATA-driven promoters are
units a, b, and b′. Three of the smaller subunits lack that also present in archaea, a key piece of evidence supporting the
relationship but are also found in RNA polymerases II and idea that in some ways, archaea resemble eukaryotes more
III. The RNA polymerases of mitochondria and chloroplasts closely than they resemble bacteria (see Table 4-1, page 102).
resemble their bacterial counterparts closely, as you might By itself, a core promoter (TATA-driven or DPE-driven) is
expect from the probable origins of these organelles as en- capable of supporting only a basal (low) level of transcription.
dosymbiotic bacteria (see Figure 4-16). Like bacterial RNA However, most protein-coding genes have additional short se-
polymerase, the mitochondrial and chloroplast enzymes are quences further upstream—upstream control elements—that
resistant to a-amanitin. improve the promoter’s efficiency. Some of these upstream
541
M18_HARD7694_09_GE_C18.indd 541 17/02/17 7:24 am
Upstream control element Core promoter
Transcriptional start site
-180 -107 -45 +1 +20
DNA Transcription
(a) Promoter for RNA polymerase I
BRE TATA box Inr DPE
Transcriptional start site
Coding-strand sequences: (G/C)(G/C)(G/A)CGCC T A T A A A A Py2 C A Py5 G(A/T)CG
-25 +1
DNA Transcription
(b) Core promoter elements for RNA polymerase II
Promoter for tRNA gene
Transcriptional start site
+1 Box A Box B
DNA Transcription
Promoter for 5S-rRNA gene
Transcriptional start site
+1 Box A Box C
DNA Transcription
(c) Two types of promoters for RNA polymerase III
Figure 18-16 Examples of Eukaryotic Promoters for RNA Polymerases I, II, and III. (a) The promoter
for RNA polymerase I has two parts, a core promoter surrounding the start site and an upstream control ele-
ment. After the binding of appropriate transcription factors to both parts, the RNA polymerase binds to the core
promoter. (b) The typical promoter for RNA polymerase II has a short initiator (Inr) sequence, consisting mostly
of pyrimidines (Py), combined with either a TATA box or a downstream promoter element (DPE). Promoters
containing a TATA box may also include a TFIIB recognition element (BRE) as part of the core promoter. (c)
The promoters for RNA polymerase III vary in structure, but the ones for tRNA genes and 5S-rRNA genes are
located entirely downstream of the start site, within the transcribed sequence. Boxes A, B, and C are DNA
consensus sequences, each about 10 bp long. In tRNA genes, about 30–60 bp of DNA separate boxes A and B.
In 5S-rRNA genes, about 10–30 bp separate boxes A and C.
elements are common to many different genes; examples and histone modifying enzymes to alter chromatin structure
include the CAAT box (consensus sequence GCCCAATCT and by promoting assembly of the basal transcriptional ma-
in animals and yeasts) and the GC box (consensus sequence chinery. Keep in mind that for genes transcribed using RNA
GGGCGG). The locations of these elements relative to a gene’s polymerase II, these long-range elements are often crucial
transcriptional start site vary from gene to gene. The elements for determining whether a gene is “switched on” (that is, it is
within 100–200 nucleotides of the start site are often called transcribed efficiently) or not (you will learn more about prox-
proximal control elements to distinguish them from enhancer imal control elements and enhancers in Chapter 20).
elements, which tend to be farther away and can even be lo- The sequences important in promoter activity are of-
cated downstream of the gene. When activating proteins bind ten identified by deleting specific sequences from a cloned
to enhancers, they locally change the conformation of DNA DNA molecule, which is then tested for its ability to serve as
near promoters by allowing chromatin remodeling proteins a template for gene transcription, either in a test tube or after
542
M18_HARD7694_09_GE_C18.indd 542 17/02/17 7:24 am
introduction of the DNA into cultured cells. For example, Core promoter
when transcription of the gene for b-globin (the b chain of Start
TATA
hemoglobin) is investigated in this way, deletion of either the DNA
TATA box or an upstream CAAT box reduces the rate of tran-
scription at least tenfold. 1 TFIID binds to TBP
In contrast to RNA polymerases I and II, the RNA poly- TATA box in DNA. TFIID
merase III molecule uses promoters that are entirely down-
stream of the transcription unit’s transcriptional start site
when transcribing genes for tRNAs and 5S rRNA. The pro-
moters used by tRNA and 5S-rRNA genes are different, but
in both cases the consensus sequences fall into two blocks of
TFIIA
about 10 bp each (Figure 18-16c). The tRNA promoter has TFIIB
consensus sequences called box A and box B. The promoters 2 TFIIA and TFIIB
form complex with
for 5S-rRNA genes have box A (positioned farther from the
TFIID.
start site than in tRNA-gene promoters) and another critical
sequence, called box C. (Not shown in the figure is a third
type of RNA polymerase III promoter, an upstream promoter
TFIIF RNA polymerase II
that is used for the synthesis of other kinds of small RNA
molecules.) C-terminal
domain
The promoters used by all the eukaryotic RNA polymer- 3 Resulting complex is
(CTD) “tail”
ases must be recognized and bound by transcription factors bound by RNA polymerase
attached to TFIIF.
before the RNA polymerase molecule can bind to DNA. We
turn now to these transcription factors.
Chapter 18
General Transcription Factors Are Involved in
the Transcription of All Nuclear Genes
|
A general transcription factor is a protein that is always
Gene Expression: I. The Genetic Code and Transcription
required for an RNA polymerase molecule to bind to its pro-
moter and initiate RNA synthesis, regardless of the identity of
the gene involved. Eukaryotes have many such transcription TFIIE
factors; their names usually include “TF” (for transcription 4 Preinitiation complex
factor), a roman numeral identifying the polymerase they aid, TFIIH is completed by addition
of TFIIE and TFIIH.
and a capital letter that identifies each individual factor (for
example, TFIIA, TFIIB).
Using RNA polymerase II as an example, Figure 18-17
illustrates the involvement of general transcription factors
in the binding of RNA polymerase to a TATA-containing pro-
moter site in DNA. General transcription factors bind to pro-
moters in a defined order, starting with TFIID. Notice that al-
though TFIID binds directly to a DNA sequence (the TATA box
ATP
in this example or the DPE sequence in the case of DPE-driven
promoters), the other transcription factors interact primarily
with each other. Hence, protein-protein interactions play a
crucial role in the binding stage of eukaryotic transcription.
RNA polymerase II does not bind to the DNA until several P
steps into the process. Eventually, a large complex of proteins, 5 RNA polymerase P P P P
CTD undergoes
including RNA polymerase, becomes bound to the promoter phosphorylation. Transcription begins
region to form a preinitiation complex.
Before RNA polymerase II can actually initiate RNA syn-
thesis, it must be released from the preinitiation complex. A Figure 18-17 Role of General Transcription Factors in
key role in this process is played by the general transcription Binding RNA Polymerase II to DNA. This figure outlines the
sequential binding of six general transcription factors (called
factor TFIIH, which possesses both a helicase activity that
TFII_, where _ is a letter identifying the particular factor) and RNA
unwinds DNA and a protein kinase activity that catalyzes
polymerase. After the final activation step involving ATP-dependent
the phosphorylation of RNA polymerase II. Phosphorylation
phosphorylation of the RNA polymerase, the polymerase can initiate
allows RNA polymerase to detach from the transcription fac- transcription. In intact chromatin, the efficient binding of general
tors so that it can initiate RNA synthesis at the transcriptional transcription factors and RNA polymerase to DNA requires the
start site. At the same time, the helicase activity of TFIIH is participation of additional regulatory proteins that open up chroma-
thought to unwind the DNA so that the RNA polymerase mol- tin structure and facilitate assembly of the preinitiation complex at
ecule can begin to move. specific genes (see Figure 20-24).
543
M18_HARD7694_09_GE_C18.indd 543 17/02/17 7:24 am
The region of RNA polymerase II that is phosphorylated lacking a TATA box, including promoters used by RNA poly-
by TFIIH is very important for regulating RNA polymerase merases I and III. Depending on the type of promoter, TBP
II function. The large subunit of RNA polymerase II has a associates with different proteins, and for promoters lacking a
C-terminal domain (CTD) that functions as a “tail.” The CTD TATA box, much of TBP’s specificity is probably derived from
is much longer than the core portion of the polymerase; it is its interaction with these associated proteins. TBP binds the
the region of the polymerase that is phosphorylated to allow minor groove of DNA, causing a severe kink in the DNA that
elongation of the RNA. The CTD contains a series of seven promotes the attachment of other components of the preini-
repeated amino acids; in yeast there are 27 of these repeats, tiation complex.
whereas in humans there are 52. Each repeat contains sites In addition to general transcription factors and RNA
that can be phosphorylated by specific kinases, including the polymerase II, several other kinds of proteins are required for
kinase that is part of TFIIH. the efficient transcription and regulated activation of specific
TFIID, the initial transcription factor to bind to the pro- genes. Some of these proteins are involved in opening up chro-
moter, is worthy of special note. Its ability to recognize and matin structure to facilitate the binding of RNA polymerase to
bind to DNA promoter sequences is conferred by one of its DNA. Others are regulatory transcription factors, which activate
subunits, the TATA-binding protein (TBP), which com- specific genes by binding to upstream control elements and
bines with a variable number of additional protein subunits recruiting coactivator proteins that in turn facilitate assembly
to form TFIID. Despite its name, the ability of TBP to bind of the RNA polymerase preinitiation complex. (The identi-
to DNA, illustrated in Figure 18-18, is not restricted to TA- ties and roles of these additional proteins will be described in
TA-containing promoters. TBP can also bind to promoters Chapter 20, which covers the regulation of gene expression;
see Figure 20-24.)
Elongation, Termination, and RNA Cleavage
Are Involved in Completing Eukaryotic RNA
Synthesis
After initiating transcription, RNA polymerases move along
the DNA and synthesize a complementary RNA copy of the
DNA template strand. Special proteins facilitate the disassem-
bly of nucleosomes in front of the moving polymerase and
their immediate reassembly after the enzyme passes. If an area
of DNA damage is encountered, RNA polymerase may become
stalled temporarily while the damage is corrected by proteins
that carry out DNA excision repair (see Figure 17-26).
TBP saddles Termination of transcription is governed by an assort-
DNA ment of signals that differ for each type of RNA polymerase.
For example, transcription by RNA polymerase I is terminated
(a) End view by a protein factor that recognizes an 18-nucleotide termina-
tion signal in the growing RNA chain. Termination signals
for RNA polymerase III are also known; they always include
TBP saddles
a short run of U’s (as in bacterial termination signals), and
DNA no ancillary protein factors are needed for their recognition.
Hairpin structures do not appear to be involved in termination
by either polymerase I or polymerase III.
For RNA polymerase II, transcripts destined to become
mRNA are often cleaved at a specific site before transcription
Kink in DNA is actually terminated. The cleavage site is 10–35 nucleotides
downstream from a special AAUAAA sequence in the grow-
ing RNA chain. The polymerase may continue transcription
for hundreds or even thousands of nucleotides beyond the
cleavage site, but this additional RNA is quickly degraded. The
cleavage site is also the site for the addition of a poly(A) tail, a
(b) Side view
string of adenine nucleotides found at the 3′ end of almost all
Figure18-18 TATA-Binding Protein (TBP) Bound to DNA. In eukaryotic mRNAs. Addition of the poly(A) tail is part of RNA
this computer graphic model, human TBP (purple) is shown bound processing, our next topic.
to DNA (blue). TBP differs from most DNA-binding proteins in that
it interacts with the minor groove of DNA, rather than the major
CONCEPT CHECK 18-2
groove, and imparts a sharp bend to the DNA. When TBP is bound
to DNA, other transcription factors can interact with the convex Compare and contrast bacterial and eukaryotic transcription,
surface of the TBP “saddle.” TBP is involved in transcription initiation focusing on initiation and termination. What features are
for all types of eukaryotic promoters. similar? What features are different?
544
M18_HARD7694_09_GE_C18.indd 544 17/02/17 7:24 am
También podría gustarte
- Acceso a Universidad para Mayores de 25 años. Biología 2013-2017.: Solucionario Pruebas 2013-2017De EverandAcceso a Universidad para Mayores de 25 años. Biología 2013-2017.: Solucionario Pruebas 2013-2017Aún no hay calificaciones
- Introducción a la Biología: RESÚMENES UNIVERSITARIOSDe EverandIntroducción a la Biología: RESÚMENES UNIVERSITARIOSCalificación: 5 de 5 estrellas5/5 (1)
- Wuolah Free TEMA 5Documento8 páginasWuolah Free TEMA 5Sara AlonsoAún no hay calificaciones
- Replicación de ADNDocumento33 páginasReplicación de ADNCarlos Alain Cabrales RamirezAún no hay calificaciones
- Clase 11 TranscripciónDocumento47 páginasClase 11 TranscripciónMARIANA MOLINA JUAREZAún no hay calificaciones
- Transcripción y Traducción en Organismos ProcariotasDocumento2 páginasTranscripción y Traducción en Organismos ProcariotasLuis Mario MartinezAún no hay calificaciones
- Sintesis, Procesamiento y Modificacion ArnDocumento59 páginasSintesis, Procesamiento y Modificacion ArnCristopher Pinedo CarranzaAún no hay calificaciones
- 12-Traducción-Fabricación de Proteínas 2020Documento31 páginas12-Traducción-Fabricación de Proteínas 2020Daniela RodriguezAún no hay calificaciones
- TranscripciónDocumento72 páginasTranscripciónNatasha PozoAún no hay calificaciones
- Transcripcion y Traduccion Del AdnDocumento39 páginasTranscripcion y Traduccion Del Adnanahi osuna100% (1)
- Dogma CentralDocumento64 páginasDogma CentralNally JumboAún no hay calificaciones
- Tipos de ArnDocumento5 páginasTipos de ArnMiguel Angel Rodas Herrera100% (2)
- Unidad 3 TranscripcionDocumento31 páginasUnidad 3 TranscripcionAngelAún no hay calificaciones
- Transcripción PDocumento25 páginasTranscripción PSHIRLEY MAYERLY CONTRERAS JAIMESAún no hay calificaciones
- Regulacion Expresión GeneticaDocumento40 páginasRegulacion Expresión GeneticaReyes Nava AliciaAún no hay calificaciones
- Transcripcion Del AdnDocumento19 páginasTranscripcion Del AdnSayda Alisson Huacre TuctoAún no hay calificaciones
- Transcripcion y TraduccionDocumento64 páginasTranscripcion y TraduccionFABIANA ANDREA DAZA ROCHAAún no hay calificaciones
- 2 Parcial de Genã©ticaDocumento17 páginas2 Parcial de Genã©ticarey reyesAún no hay calificaciones
- TranscripcionDocumento47 páginasTranscripcionFABIANA ANDREA DAZA ROCHAAún no hay calificaciones
- Clase 5 Del ADN A Las ProteinasDocumento116 páginasClase 5 Del ADN A Las ProteinasBenito VegaAún no hay calificaciones
- Mapas TranDocumento3 páginasMapas TranWilliam de Jesús López SánchezAún no hay calificaciones
- Repetición Expo. TranscripciónDocumento25 páginasRepetición Expo. TranscripciónEtyamadnessAún no hay calificaciones
- Rna InfografíaDocumento1 páginaRna InfografíaJennifer NaranjoAún no hay calificaciones
- Clase13-BiolMol-Generalidades deTranscripción-TranscProcariontesDocumento31 páginasClase13-BiolMol-Generalidades deTranscripción-TranscProcariontesJesusMondragonAún no hay calificaciones
- TranscripciónDocumento40 páginasTranscripciónfernanda lemusAún no hay calificaciones
- Transcripción Del ADN1Documento34 páginasTranscripción Del ADN1July AguilaAún no hay calificaciones
- Expresión y Regulación GénicaDocumento4 páginasExpresión y Regulación GénicaAngie Flores100% (1)
- 2) Bacteriofago T4 Lisis y LisogeniaDocumento21 páginas2) Bacteriofago T4 Lisis y LisogeniaJulian BarreraAún no hay calificaciones
- Cap 8Documento52 páginasCap 8Estefi SolangeAún no hay calificaciones
- Clase 5 y 6Documento64 páginasClase 5 y 6MatíasAún no hay calificaciones
- Estructura y Expresión Génica Eucariota IDocumento13 páginasEstructura y Expresión Génica Eucariota ILuna LorenzoAún no hay calificaciones
- 02b Transcripción en EucariontesDocumento8 páginas02b Transcripción en EucariontesMaFe CamachoAún no hay calificaciones
- BC 2021 Seminario 4 Regulacion de La Expresion de GenesDocumento59 páginasBC 2021 Seminario 4 Regulacion de La Expresion de GenesEmilio Sáez ValdésAún no hay calificaciones
- $traducción ABDocumento64 páginas$traducción ABKaren OrtizAún no hay calificaciones
- GeneticaDocumento48 páginasGeneticafernanda lemusAún no hay calificaciones
- Emma-Bioquímica-Tarea #8Documento5 páginasEmma-Bioquímica-Tarea #8Waldina LopezAún no hay calificaciones
- TranscripcionDocumento3 páginasTranscripcionNatalia VillamizarAún no hay calificaciones
- Regulación de La Transcripción y Traducción BMO42-2 2021-2 para EvaluarDocumento61 páginasRegulación de La Transcripción y Traducción BMO42-2 2021-2 para EvaluarKatya JaramilloAún no hay calificaciones
- Bioquímica I - Sem-03 - Sesion-05 - 2023-2Documento49 páginasBioquímica I - Sem-03 - Sesion-05 - 2023-2Juan AntonioAún no hay calificaciones
- Presentación Tipos de ArnDocumento11 páginasPresentación Tipos de ArnMiguel Angel Rodas HerreraAún no hay calificaciones
- 3 Dogma Central de La Biología Molecular-2010Documento73 páginas3 Dogma Central de La Biología Molecular-2010lalomestanzaAún no hay calificaciones
- Operon Lac AndresDocumento40 páginasOperon Lac AndresMiriam AlcaideAún no hay calificaciones
- Transcripción 1Documento37 páginasTranscripción 1Javiera ArellanoAún no hay calificaciones
- Transcripción PDFDocumento57 páginasTranscripción PDFManoel GallegoAún no hay calificaciones
- Expresión GénicaDocumento2 páginasExpresión GénicaAndreaAún no hay calificaciones
- Bioquímica I - Sem-03 - Sesion-05 - 2023-1Documento47 páginasBioquímica I - Sem-03 - Sesion-05 - 2023-1ANGIE REGALADO NINAQUISPEAún no hay calificaciones
- Metabolismo de ProteínasDocumento36 páginasMetabolismo de ProteínasAlberto SernaAún no hay calificaciones
- Arn Todo PDFDocumento26 páginasArn Todo PDFAnthony Alexander Marimón DiazAún no hay calificaciones
- Resumen BiocelDocumento24 páginasResumen Biocelsofiaoses1811Aún no hay calificaciones
- El Proceso de La TranscripciónDocumento46 páginasEl Proceso de La TranscripciónRosario SaldivarAún no hay calificaciones
- ARN Codificante y NO CodificanteDocumento18 páginasARN Codificante y NO CodificanteLevi OrtizAún no hay calificaciones
- Wuolah-Premium-Tema 6 - La Traducción PDFDocumento24 páginasWuolah-Premium-Tema 6 - La Traducción PDFISALEAún no hay calificaciones
- Copia de MEZA INGA Grace MarshallDocumento2 páginasCopia de MEZA INGA Grace MarshallJHADIRA ANGGIE RISALVE CUEVAAún no hay calificaciones
- C5. ARN y TranscripciónDocumento3 páginasC5. ARN y TranscripciónELISA TABOADAAún no hay calificaciones
- Biosintesis de RNA LB-Bioq Parte IDocumento17 páginasBiosintesis de RNA LB-Bioq Parte IlizetteAún no hay calificaciones
- Estructura Del Gen EucariotaDocumento76 páginasEstructura Del Gen EucariotaJuan Fabricio GravinaAún no hay calificaciones
- Organelos MembranososDocumento53 páginasOrganelos MembranososHilda SosaAún no hay calificaciones
- 02a Transcripción en ProcariontesDocumento9 páginas02a Transcripción en ProcariontesMaFe CamachoAún no hay calificaciones
- # Transcripción, Traducción y Código GenéticoDocumento18 páginas# Transcripción, Traducción y Código GenéticoJose EnriqueAún no hay calificaciones
- Diferencias EnzimáticasDocumento3 páginasDiferencias EnzimáticasDario FrancoAún no hay calificaciones