Documentos de Académico
Documentos de Profesional
Documentos de Cultura
Artificial Neural Networks - Lab 5 Self-Organising Feature Maps and Hopfield Networks
Cargado por
Ganesh Kumar ArumugamDescripción original:
Título original
Derechos de autor
Formatos disponibles
Compartir este documento
Compartir o incrustar documentos
¿Le pareció útil este documento?
¿Este contenido es inapropiado?
Denunciar este documentoCopyright:
Formatos disponibles
Artificial Neural Networks - Lab 5 Self-Organising Feature Maps and Hopfield Networks
Cargado por
Ganesh Kumar ArumugamCopyright:
Formatos disponibles
Articial Neural Networks Lab 5 Self-organising feature maps and Hopeld networks
Purpose
To study clustering with a self-organising feature map and pattern association with a Hopeld network.
Presentation
The results from the exercise should be presented individually to one of the lab organizers. If a question or sub-task appears with a box 1. then your respective answer must be presented. Students must be present at the next lab to answer questions on their work (the deadline for completion is the start of the next lab). If two students work together in a pair, then both students must be present and able to demonstrate the software that they have written and explain the answers.
Data and Matlab functions
Data and m-les can be downloaded from the course homepage (http://www.aass.oru.se/~tdt/ann). The les you need for this lab are noisy_pattern.m show_numbers.m train_hopfield.m numbers.m show_number.m test_hopfield.m test_hopfield5.m somtoolbox2.zip (contains lots of files)
In the rst task, we will use an extra toolbox for the self-organising feature map (SOFM or SOM) provided by Helsinki University of Technology (from the research lab of Prof. Teuvo Kohonen), because its a lot better than the SOFM functions provided in the Matlab neural network toolbox. For further details, take a look at http://www.cis.hut.fi/projects/somtoolbox
Preparation
Read chapters 3 and 4 of the course textbook. Note: Read each task carefully before starting to solve it.
Task 1, Learning a taxonomy with the self-organising feature map
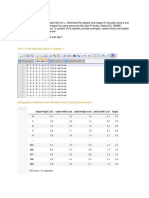
In this task, you will use Kohonens algorithm to nd clusters in data. Often you can see clusters in the 1-d or 2-d output space which you could not see in the high dimensional data simply because you cannot look at data in more than 2 dimensions (of course you can also see clusters which are spurious). To see how data can be clustered, cluster the following animals: dove, hen, duck, goose, owl, hawk, eagle, penguin, fox, dog, wolf, cat, tiger, lion, horse, zebra, cow, sheep, and pig based on the follow attributes: size:(small, medium,large); characteristics:(2 legs, 4 legs, hair, hooves, mane, feathers); behaviour:(hunt, swim, y, run) using Kohonens algorithm. Does the algorithm organize the data in a useful way? To understand how to use the SOM toolbox, here is a simple example of how to train and visualize a SOM. The example data is the well-known Iris data (it is often used as a benchmark for pattern classication systems). This data consists of 150 Iris owers divided into 3 classes: 50 Iris-setosa, 50
Articial Neural Networks Lab 5
Iris-versicolor and 50 Iris-virginica. There are four features (sepal length, sepal width, petal length, petal width), based on actual measurements taken from the owers. Look at the data le iris.data, and try to understand the le format. With the following commands you can load the data, train the SOM and label the output units of the SOM with the class names: % make the data sD = som_read_data(iris.data); sD = som_normalize(sD, var); % make the SOM sM = som_make(sD); sM = som_autolabel(sM, sD, vote); Then the SOM can be visualised using the following commands: % basic visualisation som_show(sM, umat, all, comp, 1:4, empty, Labels, norm, d); som_show_add(label, sM, subplot, 6); % plot the labels in a separate window figure som_show(sM, empty, Labels, norm, d); som_show_add(label, sM); You should see six dierent visualisations of the trained SOM: The unied distance matrix (U-matrix) visualises distances between neighbouring cluster units, and helps us to see clusters in the data. High values of the U-matrix (red cells) indicate a cluster border, while areas of low values (blue cells) indicate the clusters themselves. The next four visualisations correspond to the learned weight values for the four dierent features. For example, if you look at the rst one (feature SepalL) you should see high weight values (red) in some areas of the SOM, and low weight values (blue) in other areas. Finally, you will see the class labels that you just added to the cluster units. You can also see the labels on the cluster units, and test the SOM by nding the best matching units for all 150 owers in the data set using: % see labels on the SOM sM.labels % see SOM best matching units som_bmus(sM, sD) Use help for more details on how to use these commands. To complete this task, you should do the following: 1. Run the SOM toolbox demo som_demo1 and try to understand how it works (it will be in the exam!). Then go through the Iris example, as described above. 2. Design a suitable input coding and create your animals using the same le format as the Iris data. Show this to the lab organizer. 3. Run the training, testing and visualisation functions. Explain the results.
Articial Neural Networks Lab 5
4. Experiment with a couple of dierent network architectures, and compare with the results of 3. For example, you can create a 1-D map using something like this: sM = som_make(sD,msize,[1 10],lattice,rect); Your conclusions?
Task 2, Storage capacity of the Hopeld network
In this task, you will investigate the ability of a Hopeld network to store and retrieve patterns. The patterns that we will use are shown on page 43 of the course textbook - they are images of the numbers from 0 to 9. You can create the images by running the function numbers, then plot one image using show_number(one); or several images using show_numbers({one two three}); or show_numbers(digits); to display all of them. To help you in this task, we have implemented the training algorithm given in chapter 4 of the course textbook. You can train a Hopeld network to store two numbers using w = train_hopfield({three seven}); where w species the weight vector of the trained network. Then you can test the ability of the network to recall one of the stored patterns using x = three; x = test_hopfield(w, x, 1); where x is the pattern for recall, and 1 is the number of complete iterations of testing. You can also use test_hopfield5(w, x); to carry out 5 complete iterations of testing and display the results. Have a look at the functions, and try to work out what they do. To complete this task, you should do the following: 1. Store two patterns in a Hopeld network, and test the ability of the network to recall the same two patterns. 2. Now test the ability of the network to recall noisy versions of the same patterns. You can create noisy patterns using x = noisy_pattern(three,0.2); where the second input parameter species the amount of noise to be added to the patterns.
Articial Neural Networks Lab 5
3. Try storing three or more patterns in a Hopeld network, then test the ability of the network to recall both clean and noisy versions of the same patterns. Compare with the results of 1 and 2. Your conclusions?
4. Calculate the theoretical number of patterns that could be stored by the Hopeld network in this task, using the equation given on page 90 of the course textbook. Compare with the results of 1, 2 and 3. Your conclusions?
Namn: Godknd: a
Personnummer: Datum:
También podría gustarte
- MATLAB Machine Learning Recipes: A Problem-Solution ApproachDe EverandMATLAB Machine Learning Recipes: A Problem-Solution ApproachAún no hay calificaciones
- Lab 4 - Unsupervised Learning: K-Means ClusteringDocumento7 páginasLab 4 - Unsupervised Learning: K-Means ClusteringFaisal zafarAún no hay calificaciones
- Data Structures and Algorithms in Swift: Implement Stacks, Queues, Dictionaries, and Lists in Your AppsDe EverandData Structures and Algorithms in Swift: Implement Stacks, Queues, Dictionaries, and Lists in Your AppsAún no hay calificaciones
- Homework 3: Neural Networks and Face ImagesDocumento13 páginasHomework 3: Neural Networks and Face ImagesawesomegeeteshAún no hay calificaciones
- Laboratorio 1Documento12 páginasLaboratorio 1Daniel Israel Hidalgo RuedaAún no hay calificaciones
- Introduction To Deep Learning Assignment 0: September 2023Documento3 páginasIntroduction To Deep Learning Assignment 0: September 2023christiaanbergsma03Aún no hay calificaciones
- Week - 5 (Deep Learning) Q. 1) Explain The Architecture of Feed Forward Neural Network or Multilayer Perceptron. (12 Marks)Documento7 páginasWeek - 5 (Deep Learning) Q. 1) Explain The Architecture of Feed Forward Neural Network or Multilayer Perceptron. (12 Marks)Mrunal BhilareAún no hay calificaciones
- Advanced Coding Class PythonDocumento2 páginasAdvanced Coding Class PythonzenklerjpAún no hay calificaciones
- 11 Different Ways For Outlier Detection in PythonDocumento11 páginas11 Different Ways For Outlier Detection in PythonNeethu Merlin AlanAún no hay calificaciones
- A Neural Network Model Using PythonDocumento10 páginasA Neural Network Model Using PythonKarol SkowronskiAún no hay calificaciones
- Transform Raw Texts Into Training and Development Data: Instructor: Nikos AletrasDocumento2 páginasTransform Raw Texts Into Training and Development Data: Instructor: Nikos AletrasMD. Naimul Isalm ShovonAún no hay calificaciones
- GloveDocumento10 páginasGlovetareqeee15100% (1)
- RANDOM FOREST (Binary Classification)Documento5 páginasRANDOM FOREST (Binary Classification)Noor Ul HaqAún no hay calificaciones
- Week 2 Project - Search Algorithms - CSMMDocumento8 páginasWeek 2 Project - Search Algorithms - CSMMRaj kumar manepallyAún no hay calificaciones
- Mca1002 - Problem Solving Using Data Structures and Algorithms - LTP - 1.0 - 1 - Mca1002Documento4 páginasMca1002 - Problem Solving Using Data Structures and Algorithms - LTP - 1.0 - 1 - Mca1002Prateek BalchandaniAún no hay calificaciones
- Import OsDocumento2 páginasImport OsSumit KumarAún no hay calificaciones
- ML Unit-5Documento8 páginasML Unit-5Supriya alluriAún no hay calificaciones
- MP 3Documento3 páginasMP 3VienNgocQuangAún no hay calificaciones
- Homework: Attention-Based End-to-End Speech-to-Text Deep Neural NetworkDocumento10 páginasHomework: Attention-Based End-to-End Speech-to-Text Deep Neural NetworkAkansha KalraAún no hay calificaciones
- Big Data Machine Learning Lab 4Documento7 páginasBig Data Machine Learning Lab 4fahim.samady2001Aún no hay calificaciones
- Laboratory Work Generating A Discrete Random VariableDocumento6 páginasLaboratory Work Generating A Discrete Random VariableSufyan NikaninAún no hay calificaciones
- Deep Learning Algorithms Report PDFDocumento11 páginasDeep Learning Algorithms Report PDFrohillaanshul12Aún no hay calificaciones
- Fundamentals of Machine Learning Support Vector Machines, Practical SessionDocumento4 páginasFundamentals of Machine Learning Support Vector Machines, Practical SessionvothiquynhyenAún no hay calificaciones
- FAQ's Week-2-1Documento10 páginasFAQ's Week-2-1CharuAún no hay calificaciones
- Isolationforest1 PythonDocumento7 páginasIsolationforest1 Pythonjuan antonio garciaAún no hay calificaciones
- Homework 1Documento2 páginasHomework 1Hendra LimAún no hay calificaciones
- Step by Step TutorialDocumento8 páginasStep by Step TutorialkPrasad8Aún no hay calificaciones
- DataCamp - TensorBoard TutorialDocumento31 páginasDataCamp - TensorBoard TutorialStig KalmoAún no hay calificaciones
- Printed and Handwritten Digits Recognition Using Neural NetworksDocumento5 páginasPrinted and Handwritten Digits Recognition Using Neural NetworksParshu RamAún no hay calificaciones
- ML Lab 04 Manual - Pandas and MatplotLibDocumento7 páginasML Lab 04 Manual - Pandas and MatplotLibdodela6303Aún no hay calificaciones
- Project - Machine Learning-Business Report: By: K Ravi Kumar PGP-Data Science and Business Analytics (PGPDSBA.O.MAR23.A)Documento38 páginasProject - Machine Learning-Business Report: By: K Ravi Kumar PGP-Data Science and Business Analytics (PGPDSBA.O.MAR23.A)Ravi KotharuAún no hay calificaciones
- The Alchemy TutorialDocumento18 páginasThe Alchemy TutorialdindimieiAún no hay calificaciones
- NN Lab2Documento5 páginasNN Lab2Anne WanningenAún no hay calificaciones
- HW 1Documento3 páginasHW 1postscriptAún no hay calificaciones
- DL Mannual For ReferenceDocumento58 páginasDL Mannual For ReferenceDevant PajgadeAún no hay calificaciones
- Maxbox Starter60 Machine LearningDocumento8 páginasMaxbox Starter60 Machine LearningMax KleinerAún no hay calificaciones
- CS 201, Spring 2021: Homework Assignment 2Documento4 páginasCS 201, Spring 2021: Homework Assignment 2Arda Barış ÖrtlekAún no hay calificaciones
- STA1007S Lab 4: Scatterplots and Basic Programming: "Hist"Documento9 páginasSTA1007S Lab 4: Scatterplots and Basic Programming: "Hist"mlunguAún no hay calificaciones
- Lab 3: Unix and LinuxDocumento3 páginasLab 3: Unix and LinuxMichael FiorelliAún no hay calificaciones
- CSL465/603 - Machine LearningDocumento3 páginasCSL465/603 - Machine LearningAakarshan GuptaAún no hay calificaciones
- FML Lab 1Documento4 páginasFML Lab 1Oumaima ZiatAún no hay calificaciones
- Week3 StatDocumento4 páginasWeek3 Statsureshroyals02Aún no hay calificaciones
- DL - Assignment 1Documento12 páginasDL - Assignment 1msds21024Aún no hay calificaciones
- MIT6 189IAP11 hw2Documento8 páginasMIT6 189IAP11 hw2Ali AkhavanAún no hay calificaciones
- A 4Documento8 páginasA 4Michael PangAún no hay calificaciones
- Complete Glossary of Keras Neural Network Layers With CodeDocumento16 páginasComplete Glossary of Keras Neural Network Layers With CodeRadha KilariAún no hay calificaciones
- Ch14.A Simple Guide To MatplotlibDocumento16 páginasCh14.A Simple Guide To MatplotlibsamiluAún no hay calificaciones
- Data Science Tips and Tricks: 1. Running Jupyter Notebooks On A Shared Linux/Unix MachineDocumento6 páginasData Science Tips and Tricks: 1. Running Jupyter Notebooks On A Shared Linux/Unix Machineeight8888Aún no hay calificaciones
- SECTION 1: Basic Concepts and Notations, Arrays and RecursionDocumento6 páginasSECTION 1: Basic Concepts and Notations, Arrays and RecursionYT PremoneAún no hay calificaciones
- Assignment 1.1: First 10 Rows Looks Like Below in Notepad++Documento6 páginasAssignment 1.1: First 10 Rows Looks Like Below in Notepad++priyam100% (1)
- ML Lab 11 Manual - Neural Networks (Ver4)Documento8 páginasML Lab 11 Manual - Neural Networks (Ver4)dodela6303Aún no hay calificaciones
- JAVIER KMeans Clustering Jupyter NotebookDocumento7 páginasJAVIER KMeans Clustering Jupyter NotebookAlleah LayloAún no hay calificaciones
- Types of Machine Learning: Supervised Learning: The Computer Is Presented With Example Inputs and TheirDocumento50 páginasTypes of Machine Learning: Supervised Learning: The Computer Is Presented With Example Inputs and TheirJayesh GadewarAún no hay calificaciones
- Data Mining Algorithms in R - Clustering - Fuzzy Clustering - Fuzzy C-Means - Wikibooks, Open Books For An Open WorldDocumento8 páginasData Mining Algorithms in R - Clustering - Fuzzy Clustering - Fuzzy C-Means - Wikibooks, Open Books For An Open WorldSnr Kofi Agyarko AbabioAún no hay calificaciones
- MTH5120 Statistical Modelling I Tutorial 1Documento3 páginasMTH5120 Statistical Modelling I Tutorial 1lassti hmAún no hay calificaciones
- Neural NetworksDocumento37 páginasNeural NetworksNAIMUL HASAN NAIMAún no hay calificaciones
- Maxbox Starter100 Data Science StoryDocumento10 páginasMaxbox Starter100 Data Science StoryMax KleinerAún no hay calificaciones
- Data Structure & AlgorithmDocumento25 páginasData Structure & AlgorithmMELANIE LADRILLO ABALDEAún no hay calificaciones
- Python Solve CAT2Documento17 páginasPython Solve CAT2Er Sachin ShandilyaAún no hay calificaciones
- Intro 2 NetlabDocumento10 páginasIntro 2 NetlabBhaumik BhuvaAún no hay calificaciones
- Arkon USCX150 User GuideDocumento54 páginasArkon USCX150 User GuideGanesh Kumar ArumugamAún no hay calificaciones
- LCU Power Transducer FPWK-201Documento2 páginasLCU Power Transducer FPWK-201Ganesh Kumar ArumugamAún no hay calificaciones
- Microcomputer modulator-PTA303Documento6 páginasMicrocomputer modulator-PTA303Ganesh Kumar ArumugamAún no hay calificaciones
- Excitation System - MarunII-Thyricon Operation Manual FullDocumento61 páginasExcitation System - MarunII-Thyricon Operation Manual FullGanesh Kumar ArumugamAún no hay calificaciones
- Product Data Sheet: Auxiliary Contact, Acti9, iOF, 1 OC, AC/DCDocumento2 páginasProduct Data Sheet: Auxiliary Contact, Acti9, iOF, 1 OC, AC/DCGanesh Kumar ArumugamAún no hay calificaciones
- LVS User ManualDocumento17 páginasLVS User ManualGanesh Kumar ArumugamAún no hay calificaciones
- PMS-300 Rhino Hardware ManualDocumento39 páginasPMS-300 Rhino Hardware ManualGanesh Kumar ArumugamAún no hay calificaciones
- PPT-x80 Eagle User ManualDocumento23 páginasPPT-x80 Eagle User ManualGanesh Kumar ArumugamAún no hay calificaciones
- PMx-300 Rhino User ManualDocumento125 páginasPMx-300 Rhino User ManualGanesh Kumar ArumugamAún no hay calificaciones
- Operated and Maintenance Manual B Series...Documento286 páginasOperated and Maintenance Manual B Series...Ganesh Kumar Arumugam100% (2)
- Cable Fault Recognition Using Multiple Wavelet Neural NetworksDocumento6 páginasCable Fault Recognition Using Multiple Wavelet Neural NetworksGanesh Kumar ArumugamAún no hay calificaciones
- Ejsr 49 3 15Documento16 páginasEjsr 49 3 15Ganesh Kumar ArumugamAún no hay calificaciones
- Influnce PeopleDocumento6 páginasInflunce PeopleGanesh Kumar ArumugamAún no hay calificaciones
- Brushless DC Motors Are A Type of Synchronous MotorDocumento16 páginasBrushless DC Motors Are A Type of Synchronous MotorGanesh Kumar ArumugamAún no hay calificaciones
- Online Monitoring and Intelligent Diagnosis System Based On Hybrid Wireless Network For Large Complicated EquipmentDocumento5 páginasOnline Monitoring and Intelligent Diagnosis System Based On Hybrid Wireless Network For Large Complicated EquipmentGanesh Kumar ArumugamAún no hay calificaciones
- Fuzzy Based Fault Detection and Diagnosis in Pneumatic Actuator in Cement IndustryDocumento13 páginasFuzzy Based Fault Detection and Diagnosis in Pneumatic Actuator in Cement IndustryGanesh Kumar ArumugamAún no hay calificaciones
- Week02 - Bracketing MethodsDocumento24 páginasWeek02 - Bracketing MethodsInunHazwaniKhairuddin100% (2)
- 05 Task Performance 1Documento3 páginas05 Task Performance 1human beingAún no hay calificaciones
- cs261 hw3Documento1 páginacs261 hw3tony1337Aún no hay calificaciones
- DSP 1Documento129 páginasDSP 1Asif Eakball EmonAún no hay calificaciones
- Sparse MatrixDocumento13 páginasSparse Matrixstudy.ruby1568Aún no hay calificaciones
- Blake2 Hash - Nist Ir 7896Documento84 páginasBlake2 Hash - Nist Ir 7896fellbeasteAún no hay calificaciones
- 1.1 AlgorithmsDocumento68 páginas1.1 AlgorithmsJCAún no hay calificaciones
- CPT212 04 RecursiveDocumento26 páginasCPT212 04 RecursiveTAN KIAN HONAún no hay calificaciones
- KON 317E Control Systems Stability of Linear Control Systems Assoc. Prof. Dr. Tufan KumbasarDocumento23 páginasKON 317E Control Systems Stability of Linear Control Systems Assoc. Prof. Dr. Tufan KumbasarDogukan KarabeyinAún no hay calificaciones
- CSCI 2100 Data StructuresDocumento3 páginasCSCI 2100 Data Structuresnishant.vepachedu8644Aún no hay calificaciones
- FADML 07 PPC Minimum Spanning Trees PDFDocumento16 páginasFADML 07 PPC Minimum Spanning Trees PDFarpan singhAún no hay calificaciones
- 2.1 Survey On Parallel Fir FiltersDocumento19 páginas2.1 Survey On Parallel Fir FilterskishorechiyaAún no hay calificaciones
- Matrix Decomposition - WikipediaDocumento10 páginasMatrix Decomposition - WikipediaDebashish RoulAún no hay calificaciones
- hw3 19fDocumento2 páginashw3 19fAmreshAmanAún no hay calificaciones
- GP - DSA - Kadane's AlgorithmDocumento3 páginasGP - DSA - Kadane's AlgorithmOm BhosaleAún no hay calificaciones
- CS1010S-Lec-03 Recursion, IterationDocumento70 páginasCS1010S-Lec-03 Recursion, IterationAkane YoriAún no hay calificaciones
- Assignment ImageDocumento5 páginasAssignment ImageHafiz AmirulAún no hay calificaciones
- Advanced Exposure Fusion Using New Boosting Laplacian Pyramid REVIEW 2Documento15 páginasAdvanced Exposure Fusion Using New Boosting Laplacian Pyramid REVIEW 2Ramarao GudeAún no hay calificaciones
- Must Solve 100 Programs Part 2 - 10Documento11 páginasMust Solve 100 Programs Part 2 - 10Rohit RoushanAún no hay calificaciones
- Finite Difference MethodDocumento6 páginasFinite Difference MethodNovieOppy November BiAún no hay calificaciones
- 2017 Fall ME349 03 NumAnalysis1Documento41 páginas2017 Fall ME349 03 NumAnalysis1Abdo SalahAún no hay calificaciones
- 1.supervised and UnsupervisedDocumento42 páginas1.supervised and Unsupervisedrajthakre81Aún no hay calificaciones
- Signal Transmission Through A Linear System, Signal Distortion Over A Communication Channel, Measure of Signals, ESD and PSD of Modulated SignalsDocumento3 páginasSignal Transmission Through A Linear System, Signal Distortion Over A Communication Channel, Measure of Signals, ESD and PSD of Modulated SignalsCool GuyAún no hay calificaciones
- Objectives:: at The End of The Course, The Student Should Be Able ToDocumento2 páginasObjectives:: at The End of The Course, The Student Should Be Able TomakAún no hay calificaciones
- DAA U-4 Dynamic ProgrammingDocumento23 páginasDAA U-4 Dynamic Programmings.s.r.k.m. guptaAún no hay calificaciones
- Mathematics For Economics (ECON 104)Documento46 páginasMathematics For Economics (ECON 104)Experimental BeXAún no hay calificaciones
- Exploration of Reinforcement Learning To SNAKE: Bowei Ma, Meng Tang, Jun ZhangDocumento5 páginasExploration of Reinforcement Learning To SNAKE: Bowei Ma, Meng Tang, Jun ZhangNada JonidiAún no hay calificaciones
- 1 Uber Interview Experience (On Campus For Internship 2018-19) 2Documento5 páginas1 Uber Interview Experience (On Campus For Internship 2018-19) 2Ajay KumarAún no hay calificaciones
- Lecture07 SlidesDocumento42 páginasLecture07 SlidesYunyang LiAún no hay calificaciones
- Generative AI: The Insights You Need from Harvard Business ReviewDe EverandGenerative AI: The Insights You Need from Harvard Business ReviewCalificación: 4.5 de 5 estrellas4.5/5 (2)
- ChatGPT Side Hustles 2024 - Unlock the Digital Goldmine and Get AI Working for You Fast with More Than 85 Side Hustle Ideas to Boost Passive Income, Create New Cash Flow, and Get Ahead of the CurveDe EverandChatGPT Side Hustles 2024 - Unlock the Digital Goldmine and Get AI Working for You Fast with More Than 85 Side Hustle Ideas to Boost Passive Income, Create New Cash Flow, and Get Ahead of the CurveAún no hay calificaciones
- Scary Smart: The Future of Artificial Intelligence and How You Can Save Our WorldDe EverandScary Smart: The Future of Artificial Intelligence and How You Can Save Our WorldCalificación: 4.5 de 5 estrellas4.5/5 (55)
- The Digital Mind: How Science is Redefining HumanityDe EverandThe Digital Mind: How Science is Redefining HumanityCalificación: 4.5 de 5 estrellas4.5/5 (2)
- Working with AI: Real Stories of Human-Machine Collaboration (Management on the Cutting Edge)De EverandWorking with AI: Real Stories of Human-Machine Collaboration (Management on the Cutting Edge)Calificación: 5 de 5 estrellas5/5 (5)
- ChatGPT Money Machine 2024 - The Ultimate Chatbot Cheat Sheet to Go From Clueless Noob to Prompt Prodigy Fast! Complete AI Beginner’s Course to Catch the GPT Gold Rush Before It Leaves You BehindDe EverandChatGPT Money Machine 2024 - The Ultimate Chatbot Cheat Sheet to Go From Clueless Noob to Prompt Prodigy Fast! Complete AI Beginner’s Course to Catch the GPT Gold Rush Before It Leaves You BehindAún no hay calificaciones
- The Master Algorithm: How the Quest for the Ultimate Learning Machine Will Remake Our WorldDe EverandThe Master Algorithm: How the Quest for the Ultimate Learning Machine Will Remake Our WorldCalificación: 4.5 de 5 estrellas4.5/5 (107)
- Four Battlegrounds: Power in the Age of Artificial IntelligenceDe EverandFour Battlegrounds: Power in the Age of Artificial IntelligenceCalificación: 5 de 5 estrellas5/5 (5)
- ChatGPT: Is the future already here?De EverandChatGPT: Is the future already here?Calificación: 4 de 5 estrellas4/5 (21)
- 100M Offers Made Easy: Create Your Own Irresistible Offers by Turning ChatGPT into Alex HormoziDe Everand100M Offers Made Easy: Create Your Own Irresistible Offers by Turning ChatGPT into Alex HormoziAún no hay calificaciones
- HBR's 10 Must Reads on AI, Analytics, and the New Machine AgeDe EverandHBR's 10 Must Reads on AI, Analytics, and the New Machine AgeCalificación: 4.5 de 5 estrellas4.5/5 (69)
- Artificial Intelligence & Generative AI for Beginners: The Complete GuideDe EverandArtificial Intelligence & Generative AI for Beginners: The Complete GuideCalificación: 5 de 5 estrellas5/5 (1)
- The AI Advantage: How to Put the Artificial Intelligence Revolution to WorkDe EverandThe AI Advantage: How to Put the Artificial Intelligence Revolution to WorkCalificación: 4 de 5 estrellas4/5 (7)
- T-Minus AI: Humanity's Countdown to Artificial Intelligence and the New Pursuit of Global PowerDe EverandT-Minus AI: Humanity's Countdown to Artificial Intelligence and the New Pursuit of Global PowerCalificación: 4 de 5 estrellas4/5 (4)
- Python Machine Learning - Third Edition: Machine Learning and Deep Learning with Python, scikit-learn, and TensorFlow 2, 3rd EditionDe EverandPython Machine Learning - Third Edition: Machine Learning and Deep Learning with Python, scikit-learn, and TensorFlow 2, 3rd EditionCalificación: 5 de 5 estrellas5/5 (2)
- Artificial Intelligence: The Insights You Need from Harvard Business ReviewDe EverandArtificial Intelligence: The Insights You Need from Harvard Business ReviewCalificación: 4.5 de 5 estrellas4.5/5 (104)
- Your AI Survival Guide: Scraped Knees, Bruised Elbows, and Lessons Learned from Real-World AI DeploymentsDe EverandYour AI Survival Guide: Scraped Knees, Bruised Elbows, and Lessons Learned from Real-World AI DeploymentsAún no hay calificaciones
- Artificial Intelligence: A Guide for Thinking HumansDe EverandArtificial Intelligence: A Guide for Thinking HumansCalificación: 4.5 de 5 estrellas4.5/5 (30)
- Architects of Intelligence: The truth about AI from the people building itDe EverandArchitects of Intelligence: The truth about AI from the people building itCalificación: 4.5 de 5 estrellas4.5/5 (21)
- ChatGPT Millionaire 2024 - Bot-Driven Side Hustles, Prompt Engineering Shortcut Secrets, and Automated Income Streams that Print Money While You Sleep. The Ultimate Beginner’s Guide for AI BusinessDe EverandChatGPT Millionaire 2024 - Bot-Driven Side Hustles, Prompt Engineering Shortcut Secrets, and Automated Income Streams that Print Money While You Sleep. The Ultimate Beginner’s Guide for AI BusinessAún no hay calificaciones
- The Roadmap to AI Mastery: A Guide to Building and Scaling ProjectsDe EverandThe Roadmap to AI Mastery: A Guide to Building and Scaling ProjectsAún no hay calificaciones
- How To: GPT-4 for Financial Freedom: The Entrepreneur's Guide to Thriving in the AI EraDe EverandHow To: GPT-4 for Financial Freedom: The Entrepreneur's Guide to Thriving in the AI EraCalificación: 4 de 5 estrellas4/5 (9)
- Mastering Large Language Models: Advanced techniques, applications, cutting-edge methods, and top LLMs (English Edition)De EverandMastering Large Language Models: Advanced techniques, applications, cutting-edge methods, and top LLMs (English Edition)Aún no hay calificaciones
- Who's Afraid of AI?: Fear and Promise in the Age of Thinking MachinesDe EverandWho's Afraid of AI?: Fear and Promise in the Age of Thinking MachinesCalificación: 4.5 de 5 estrellas4.5/5 (13)