Documentos de Académico
Documentos de Profesional
Documentos de Cultura
Michaelis-Menten Kinetics
Cargado por
charanmann9165Título original
Derechos de autor
Formatos disponibles
Compartir este documento
Compartir o incrustar documentos
¿Le pareció útil este documento?
¿Este contenido es inapropiado?
Denunciar este documentoCopyright:
Formatos disponibles
Michaelis-Menten Kinetics
Cargado por
charanmann9165Copyright:
Formatos disponibles
MichaelisMenten kinetics
Concentration
[S]
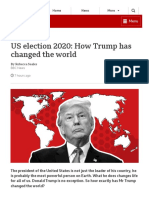
MichaelisMenten saturation curve for an enzyme reaction
showing the relation between the substrate concentration and reaction rate.
[P]
[E]
[ES]
Time
In biochemistry, MichaelisMenten kinetics is one of
the best-known models of enzyme kinetics. It is named
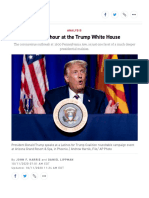
after German biochemist Leonor Michaelis and Canadian Change in concentrations over time for enzyme E, substrate S,
physician Maud Menten. The model takes the form of complex ES and product P
an equation describing the rate of enzymatic reactions,
by relating reaction rate v to [S] , the concentration of a
where kf , kr , and kcat denote the rate constants,[5] and
substrate S. Its formula is given by
the double arrows between S and ES represent the fact
that enzyme-substrate binding is a reversible process.
v=
d[P ]
Vmax [S]
=
.
dt
KM + [S]
Under certain assumptions such as the enzyme concentration being much less than the substrate concentration
the rate of product formation is given by
Here, Vmax represents the maximum rate achieved by
the system, at maximum (saturating) substrate concentrations. The Michaelis constant KM is the substrate cond[P ]
[S]
[S]
centration at which the reaction rate is half of Vmax .[1] v =
= Vmax
= kcat [E]0
.
dt
KM + [S]
KM + [S]
Biochemical reactions involving a single substrate are often assumed to follow MichaelisMenten kinetics, with- The reaction rate increases with increasing substrate conout regard to the models underlying assumptions.
centration [S] , asymptotically approaching its maximum
rate Vmax , attained when all enzyme is bound to substrate.
It also follows that Vmax = kcat [E]0 , where [E]0 is the
1 Model
initial enzyme concentration. kcat , the turnover number,
is the maximum number of substrate molecules converted
In 1903, French physical chemist Victor Henri found that to product per enzyme molecule per second.
enzyme reactions were initiated by a bond (more gen- The Michaelis constant KM is the substrate concentraerally, a binding interaction) between the enzyme and tion at which the reaction rate is at half-maximum,[1] and
the substrate.[2] His work was taken up by German bio- is an inverse measure of the substrates anity for the
chemist Leonor Michaelis and Canadian physician Maud enzymeas a small KM indicates high anity, meanMenten, who investigated the kinetics of an enzymatic re- ing that the rate will approach Vmax more quickly.[6] The
action mechanism, invertase, that catalyzes the hydrolysis value of KM is dependent on both the enzyme and the
of sucrose into glucose and fructose.[3] In 1913, they pro- substrate, as well as conditions such as temperature and
posed a mathematical model of the reaction.[4] It involves pH.
an enzyme E binding to a substrate S to form a complex
ES, which in turn is converted into a product P and the The model is used in a variety of biochemical situations other than enzyme-substrate interaction, including
enzyme. This may be represented schematically as
antigen-antibody binding, DNA-DNA hybridization, and
protein-protein interaction.[6][7] It can be used to charkf
acterise a generic biochemical reaction, in the same way
kcat
E+P
E + S ES
that the Langmuir equation can be used to model generic
kr
1
DERIVATION
adsorption of biomolecular species.[7] When an empirical
equation of this form is applied to microbial growth, it is
sometimes called a Monod equation.
[E] = [E]0 [ES].
Combining the two expressions above, gives us
Applications
Parameter values vary wildly between enzymes:[8]
kf ([E]0 [ES])[S] = kr [ES].
The constant kcat /KM (catalytic eciency) is a measure Upon simplication, we get
of how an enzyme eciently converts a substrate into
product. Diusion limited enzymes, such as fumarase,
work at the theoretical upper limit of 108 1010 /M*s,
[E]0 [S]
limited by diusion of substrate into the active site.[9]
[ES] =
Kd + [S]
MichaelisMenten kinetics have also been applied to a
variety of spheres outside of biochemical reactions,[5] where Kd = kr /kf is the dissociation constant for the
including alveolar clearance of dusts,[10] the richness enzyme-substrate complex. Hence the velocity v of the
of species pools,[11] clearance of blood alcohol,[12] reaction the rate at which P is formed is[15]
the photosynthesis-irradiance relationship, and bacterial
phage infection.[13]
v=
Derivation
d[P ]
Vmax [S]
=
dt
Kd + [S]
where Vmax = kcat [E]0 is the maximum reaction velocity.
Applying the law of mass action, which states that the rate
of a reaction is proportional to the product of the concentrations of the reactants (i.e.[E][S]), gives a system of 3.2 Quasi-steady-state approximation
four non-linear ordinary dierential equations that dene
An alternative analysis of the system was undertaken
the rate of change of reactants with time t [14]
by British botanist G. E. Briggs and British geneticist J. B. S. Haldane in 1925.[16] They assumed that
the concentration of the intermediate complex does not
d[E]
= kf [E][S] + kr [ES] + kcat [ES]
change on the time-scale of product formation known
dt
as the quasi-steady-state assumption or pseudo-steadyd[S]
= kf [E][S] + kr [ES]
state-hypothesis. Mathematically, this assumption means
dt
kf [E][S] = kr [ES] + kcat [ES] . Combining this relad[ES]
= kf [E][S] kr [ES] kcat [ES]
tionship with the enzyme conservation law, the concendt
tration of complex is[15]
d[P ]
= kcat [ES].
dt
In this mechanism, the enzyme E is a catalyst, which only
[E]0 [S]
[ES] =
facilitates the reaction, so that its total concentration, free
KM + [S]
plus combined, [E] + [ES] = [E]0 is a constant. This
conservation law can also be observed by adding the rst where
and third equations above.[14][15]
3.1
Equilibrium approximation
KM =
kr + kcat
kf
In their original analysis, Michaelis and Menten assumed is known as the Michaelis constant, where kr , kcat , and
that the substrate is in instantaneous chemical equilibrium kf are, respectively, the constants for substrate unbinding,
with the complex, which implies[4][15]
conversion to product, and binding to the enzyme. Hence
the velocity v of the reaction is[15]
kf [E][S] = kr [ES].
From the enzyme conservation law, we obtain[15]
v=
d[P ]
Vmax [S]
=
.
dt
KM + [S]
3.3
Assumptions and limitations
The rst step in the derivation applies the law of mass action, which is reliant on free diusion. However, in the
environment of a living cell where there is a high concentration of proteins, the cytoplasm often behaves more
like a gel than a liquid, limiting molecular movements and
altering reaction rates.[17] Whilst the law of mass action
can be valid in heterogeneous environments,[18] it is more
appropriate to model the cytoplasm as a fractal, in order
to capture its limited-mobility kinetics.[19]
[S] [P ].
This is true under standard in vitro assay conditions, and
is true for many in vivo biological reactions, particularly
where the product is continually removed by a subsequent
reaction.
2. The energy released in the reaction is very
large, that is
The resulting reaction rates predicted by the two approaches are similar, with the only dierence being that
the equilibrium approximation denes the constant as Kd
, whilst the quasi-steady-state approximation uses KM .
G 0.
However, each approach is founded upon a dierent assumption. The MichaelisMenten equilibrium analysis is
valid if the substrate reaches equilibrium on a much faster In situations where neither of these two conditions hold
time-scale than the product is formed or, more precisely, (that is, the reaction is low energy and a substantial pool of
that [15]
product(s) exists), the MichaelisMenten equation breaks
down, and more complex modelling approaches explicitly taking the forward and reverse reactions into account
kcat
must be taken to understand the enzyme biology.
d =
1.
kr
By contrast, the BriggsHaldane quasi-steady-state analysis is valid if [14][20]
m =
[E]0
1.
[S]0 + KM
Thus it holds if the enzyme concentration is much less
than the substrate concentration. Even if this is not satised, the approximation is valid if KM is large.
In both the MichaelisMenten and BriggsHaldane analyses, the quality of the approximation improves as decreases. However, in model building, MichaelisMenten
kinetics are often invoked without regard to the underlying assumptions.[15]
It is also important to remember that, while irreversibility
is a necessary simplication in order to yield a tractable
analytic solution, in the general case product formation
is not in fact irreversible. The enzyme reaction is more
correctly described as
kf1
kf2
kr1
kr2
E + S ES E + P.
4 Determination of constants
The typical method for determining the constants Vmax
and KM involves running a series of enzyme assays at
varying substrate concentrations [S] , and measuring the
initial reaction rate v0 . 'Initial' here is taken to mean
that the reaction rate is measured after a relatively short
time period, during which it is assumed that the enzymesubstrate complex has formed, but that the substrate
concentration held approximately constant, and so the
equilibrium or quasi-steady-state approximation remain
valid.[20] By plotting reaction rate against concentration,
and using nonlinear regression of the MichaelisMenten
equation, the parameters may be obtained.[21]
Before computing facilities to perform nonlinear regression became available, graphical methods involving linearisation of the equation were used. A number of these
were proposed, including the EadieHofstee diagram,
HanesWoolf plot and LineweaverBurk plot; of these,
the HanesWoolf plot is the most accurate.[21] However,
while useful for visualization, all three methods distort
the error structure of the data and are inferior to nonlinear regression.[22] Nonetheless, their use can still be found
in modern literature.[23]
In general, the assumption of irreversibility is a good one In 1997 Santiago Schnell and Claudio Mendoza suggested
a closed form solution for the time course kinetics analin situations where one of the below is true:
ysis of the MichaelisMenten kinetics based on the solution of the Lambert W function.[24] Namely:
1. The concentration of substrate(s) is very
much larger than the concentration of products:
[S]
= W (F (t))
KM
where W is the Lambert W function and
(
)
[S]0
[S]0
Vmax
F (t) =
exp
t .
KM
KM
KM
REFERENCES
[5] Chen, W.W.; Neipel, M.; Sorger, P.K. (2010). Classic
and contemporary approaches to modeling biochemical reactions.
Genes Dev 24 (17): 18611875.
doi:10.1101/gad.1945410. PMC 2932968. PMID
20810646.
The above equation has been used to estimate Vmax and
KM from time course data.[25][26]
[6] Lehninger, A.L.; Nelson, D.L.; Cox, M.M. (2005).
Lehninger principles of biochemistry. New York: W.H.
Freeman. ISBN 978-0-7167-4339-2.
[7] Chakraborty, S. (23 Dec 2009). Microuidics and Microfabrication (1 ed.). Springer. ISBN 978-1-4419-1542-9.
Role of substrate unbinding
[8] Mathews, C.K.; van Holde, K.E.; Ahern, K.G. (10 Dec
The Michaelis-Menten equation has been used to pre1999). Biochemistry (3 ed.). Prentice Hall. ISBN 978-0dict the rate of product formation in enzymatic reactions
8053-3066-3.
for more than a century. Specically, it states that the
rate of an enzymatic reaction will increase as substrate [9] Stroppolo, M.E.; Falconi, M.; Caccuri, A.M.; Desideri,
concentration increases, and that increased unbinding of
A. (Sep 2001). Superecient enzymes. Cell Mol Life
enzyme-substrate complexes will decrease the reaction
Sci 58 (10): 145160. doi:10.1007/PL00000788. PMID
11693526.
rate. While the rst prediction is well established, the second has never been tested experimentally. To determine
whether an increased rate of unbinding does in fact de- [10] Yu, R.C.; Rappaport, S.M. (1997). A lung retention model based on MichaelisMenten-like kinetcrease the reaction rate, Shlomi Reuveni et al. mathematics.
Environ Health Perspect 105 (5): 496503.
ically analyzed the eect of enzyme-substrate unbinddoi:10.1289/ehp.97105496. PMC 1469867. PMID
ing on enzymatic reactions at the single-molecule level.
9222134.
According to the study, unbinding of an enzyme from a
substrate can reduce the rate of product formation under [11] Keating, K.A.; Quinn, J.F. (1998). Estimating species
some conditions, but may also have the opposite eect.
richness: the MichaelisMenten model revisited. Oikos
As substrate concentrations increase, a tipping point can
81 (2): 411416. doi:10.2307/3547060. JSTOR
3547060.
be reached where an increase in the unbinding rate results
in an increase, rather than a decrease, of the reaction rate.
The results indicate that enzymatic reactions can behave [12] Jones, A.W. (2010). Evidence-based survey of the elimination rates of ethanol from blood with applications in
in ways that violate the classical Michaelis-Menten equaforensic casework. Forensic Sci Int 200 (13): 120.
tion, and that the role of unbinding in enzymatic catalysis
doi:10.1016/j.forsciint.2010.02.021. PMID 20304569.
[27]
still remains to be determined experimentally.
See also
[13] Abedon, S.T. (2009). Kinetics of phage-mediated biocontrol of bacteria. Foodborne Pathog Dis 6 (7): 80715.
doi:10.1089/fpd.2008.0242. PMID 19459758.
Enzyme kinetics
[14] Murray, J.D. (2002). Mathematical Biology: I. An Introduction (3 ed.). Springer. ISBN 978-0-387-95223-9.
LineweaverBurk plot
Reaction progress kinetic analysis
Steady state (chemistry)
References
[1] http://www.worthington-biochem.com/introbiochem/
substrateconc.html
[2] Henri, Victor (1903). Lois Gnrales de lAction des Diastases. Paris: Hermann. Google books (US only)
[3] Victor Henri. Whonamedit?. Retrieved 24 May 2011.
[4] Michaelis, L.; Menten, M.L. (1913). Die Kinetik der
Invertinwirkung. Biochem Z 49: 333369 (recent translation, and an older partial translation)
[15] Keener, J.; Sneyd, J. (2008). Mathematical Physiology: I:
Cellular Physiology (2 ed.). Springer. ISBN 978-0-38775846-6.
[16] Briggs, G.E.; Haldane, J.B.S. (1925). A note on the kinematics of enzyme action. Biochem J 19 (2): 338339.
PMC 1259181. PMID 16743508.
[17] Zhou, H.X.; Rivas, G.; Minton, A.P. (2008).
Macromolecular crowding and connement: biochemical, biophysical, and potential physiological
consequences. Annu Rev Biophys 37 (1): 37597.
doi:10.1146/annurev.biophys.37.032807.125817. PMC
2826134. PMID 18573087.
[18] Grima, R.; Schnell, S. (Oct 2006).
A systematic investigation of the rate laws valid in intracellular environments. Biophys Chem 124 (1): 110.
doi:10.1016/j.bpc.2006.04.019. PMID 16781049.
[19] Schnell, S.; Turner, T.E. (2004). Reaction kinetics in
intracellular environments with macromolecular crowding: simulations and rate laws. Prog Biophys Mol Biol 85
(23): 23560. doi:10.1016/j.pbiomolbio.2004.01.012.
PMID 15142746.
[20] Segel, L.A.; Slemrod, M. (1989). The quasi-steady-state
assumption: A case study in perturbation. Thermochim
Acta 31 (3): 446477. doi:10.1137/1031091.
[21] Leskovac, V. (2003). Comprehensive enzyme kinetics.
New York: Kluwer Academic/Plenum Pub. ISBN 9780-306-46712-7.
[22] Greco, W.R.; Hakala, M.T. (1979). Evaluation of methods for estimating the dissociation constant of tight binding enzyme inhibitors,. J Biol Chem 254 (23): 12104
12109. PMID 500698.
[23] Hayakawa, K.; Guo, L.; Terentyeva, E.A.; Li, X.K.;
Kimura, H.; Hirano, M.; Yoshikawa, K.; Nagamine,
T.; et al. (2006). Determination of specic activities and kinetic constants of biotinidase and lipoamidase
in LEW rat and Lactobacillus casei (Shirota)". J Chromatogr B Analyt Technol Biomed Life Sci 844 (2): 24050.
doi:10.1016/j.jchromb.2006.07.006. PMID 16876490.
[24] Schnell, S.; Mendoza, C. (1997). A closed form
solution for time-dependent enzyme kinetics.
Journal of Theoretical Biology 187 (2): 207212.
doi:10.1006/jtbi.1997.0425.
[25] Goudar, C. T.; Sonnad, J. R.; Duggleby, R. G. (1999).
Parameter estimation using a direct solution of the integrated MichaelisMenten equation. Biochimica et
Biophysica Acta Protein Structure and Molecular Enzymology 1429 (2): 377383. doi:10.1016/s01674838(98)00247-7. PMID 9989222.
[26] Goudar, C. T.; Harris, S. K.; McInerney, M. J.; Suita, J. M. (2004). Progress curve analysis for enzyme and microbial kinetic reactions using explicit solutions based on the Lambert W function. Journal of Microbiological Methods 59 (3): 317326.
doi:10.1016/j.mimet.2004.06.013. PMID 15488275.
[27] Reuveni, Shlomi;
Urbakh, Michael;
Klafter,
Joseph (2014).
Role of Substrate Unbinding in
Michaelis-Menten Enzymatic Reactions.
Proceedings of the National Academy of Sciences 111
(12): 43914396. Bibcode:2014PNAS..111.4391R.
doi:10.1073/pnas.1318122111.
Further reading
Biochemistry/Catalysis at Wikibooks
9 TEXT AND IMAGE SOURCES, CONTRIBUTORS, AND LICENSES
Text and image sources, contributors, and licenses
9.1
Text
MichaelisMenten kinetics Source: https://en.wikipedia.org/wiki/Michaelis%E2%80%93Menten_kinetics?oldid=691358742 Contributors: Michael Hardy, Lexor, Charles Matthews, Ike9898, Wik, Aliekens, Lord Kelvin, Cutler, Giftlite, Jao, Bensaccount, Delta G,
Christopherlin, Avihu, Thorwald, DanielCD, Rich Farmbrough, Cacycle, Bender235, Iamunknown, Lenov, Maurreen, Arcadian, Axl,
Stillnotelf, Vuo, Wsloand, Gene Nygaard, Richmc, Palica, V8rik, Rjwilmsi, John Baez, Mathbot, Matro~enwiki, Hede2000, Shell Kinney, NawlinWiki, Deskana, Jrf, Cubic Hour, Eykanal, Itub, KnightRider~enwiki, Slashme, Huhnra, David Shear, Stepa, Eskimbot, CuriousOliver, Chris the speller, Mion, SashatoBot, Sasata, Gegnome, CzarB, Wfgiuliano, Kaarel, Lvzon, CmdrObot, Pro bug catcher,
Fij, Christian75, Calvero JP, Thijs!bot, TimVickers, Isilanes, Boleslaw, Twisted86, JaGa, Squidonius, Ultraviolet scissor ame, Xris0,
SjoerdC, Hannes Rst, Northfox, SHL-at-Sv, Ian Glenn, SieBot, Maphyche, Denisarona, Deviator13, Genie05, PixelBot, Cschulthess,
Clayt85, Agor153, Tiphaine800, MystBot, Addbot, Athenray, Diptanshu.D, Download, 84user, Martina Steiner, Tide rolls, Lightbot,
Yobot, Expresser, Rudolf.hellmuth, Materialscientist, Citation bot, Eumolpo, Obersachsebot, Aa77zz, ChristopherKingChemist, Anrade,
Lostella~enwiki, Citation bot 1, Waveguide2, RC Howe, Lmp883, Rachelpurdon, Slawekb, Stevestoker, Vramasub, Shazux, Andkennard,
U+003F, Bulto95, Cornell92, Kinkreet, ClueBot NG, Smenden, Gareth Grith-Jones, SSSheridan, Lifeonahilltop, Widr, Helpful Pixie
Bot, Djmaity, BG19bot, C.Rose.Kennedy.2, Wenlanzsw, NotoriousOCG, D Scott 86, Dy195195, TristanCroll, Fkendrick3, Evolution and
evolvability, YiFeiBot, Earley11, Monkbot, Shibbolethink, Czeer, Shlomireuveni, Harryrockfellaz and Anonymous: 144
9.2
Images
File:Michaelis_Menten_S_P_E_ES.svg Source: https://upload.wikimedia.org/wikipedia/commons/2/23/Michaelis_Menten_S_P_E_
ES.svg License: CC0 Contributors: Own work Original artist: U+003F
File:Michaelis_Menten_curve_2.svg Source: https://upload.wikimedia.org/wikipedia/commons/8/83/Michaelis_Menten_curve_2.svg
License: CC BY-SA 4.0 Contributors: Own work Original artist: Thomas Shafee
File:Nuvola_apps_edu_science.svg Source: https://upload.wikimedia.org/wikipedia/commons/5/59/Nuvola_apps_edu_science.svg License: LGPL Contributors: http://ftp.gnome.org/pub/GNOME/sources/gnome-themes-extras/0.9/gnome-themes-extras-0.9.0.tar.gz Original artist: David Vignoni / ICON KING
9.3
Content license
Creative Commons Attribution-Share Alike 3.0
También podría gustarte
- Agenda Studio in AutonomiaDocumento3 páginasAgenda Studio in AutonomiafazzioAún no hay calificaciones
- Materials Analysis by Ion Channeling: Submicron CrystallographyDe EverandMaterials Analysis by Ion Channeling: Submicron CrystallographyAún no hay calificaciones
- Palestrina LamentationsDocumento29 páginasPalestrina LamentationsPaul DeleuzeAún no hay calificaciones
- The Elgar Companion of Health EconomicsDocumento584 páginasThe Elgar Companion of Health EconomicsApostolos Davillas100% (1)
- Graphs ModulationDocumento15 páginasGraphs Modulationkowskii666_719410288100% (2)
- Precis AnalysisDocumento11 páginasPrecis AnalysisFederico Favali100% (1)
- Essential Neo-Riemannian Theory For Todays MusicianDocumento98 páginasEssential Neo-Riemannian Theory For Todays MusicianNicolas FinkelsteinAún no hay calificaciones
- A Critical Edition On The 48 Studies For OBOE - Vol2 PDFDocumento180 páginasA Critical Edition On The 48 Studies For OBOE - Vol2 PDFsetipareAún no hay calificaciones
- ESMC2018 Technical Program Web 0 PDFDocumento291 páginasESMC2018 Technical Program Web 0 PDFSenthilkumar VAún no hay calificaciones
- Mediveal MusicDocumento3 páginasMediveal MusicKahfiar Anugerah MawaddahAún no hay calificaciones
- Eserciziario Di Analisi Matematica 2Documento322 páginasEserciziario Di Analisi Matematica 2Giuseppe MoreseAún no hay calificaciones
- Armonia OttatonicaDocumento18 páginasArmonia Ottatonicabart100% (1)
- Pocci Catalog 33th February 2018 Composers Profiles PDFDocumento518 páginasPocci Catalog 33th February 2018 Composers Profiles PDFAires Pinheiro100% (1)
- Ommagio A JoyceDocumento3 páginasOmmagio A JoyceDaniel KaramazovAún no hay calificaciones
- Organology I: Plant Structure and DevelopmentDocumento6 páginasOrganology I: Plant Structure and DevelopmentSuryadi SuryadiAún no hay calificaciones
- Music Theory and Analysis:: Musical Semiotics 40 Years AfterDocumento32 páginasMusic Theory and Analysis:: Musical Semiotics 40 Years AfterZoran Nastov100% (1)
- Vanscheeuwijck, Marc - Musical Performance at San Petronio in Bologna A Brief HistoryDocumento11 páginasVanscheeuwijck, Marc - Musical Performance at San Petronio in Bologna A Brief HistoryKaren BarbosaAún no hay calificaciones
- Grosse Fuge - AnalysisDocumento10 páginasGrosse Fuge - AnalysiscalusAún no hay calificaciones
- DeepBach - A Steerable Model For Bach Chorales GenerationDocumento10 páginasDeepBach - A Steerable Model For Bach Chorales GenerationFiammetta GhediniAún no hay calificaciones
- Elgar Cello Concerto and Rouse Flute Concerto: Responses To 20 Century TragedyDocumento11 páginasElgar Cello Concerto and Rouse Flute Concerto: Responses To 20 Century Tragedypigio_4946403Aún no hay calificaciones
- Chapter 15Documento8 páginasChapter 15Tilak K CAún no hay calificaciones
- Ccwatershed.org website linksDocumento137 páginasCcwatershed.org website linksFrancesco DeffenuAún no hay calificaciones
- 06 Chapter 1Documento55 páginas06 Chapter 1Steven HowellAún no hay calificaciones
- Simple Kinetics of Enzyme ActionDocumento7 páginasSimple Kinetics of Enzyme ActionRavindra Mani TiwariAún no hay calificaciones
- Michaelis MentenKineticsandBriggs HaldaneKinetics - HTMLDocumento2 páginasMichaelis MentenKineticsandBriggs HaldaneKinetics - HTMLCrestore Lex Tapia CapiñaAún no hay calificaciones
- Lecture 14: Enzyme Kinetics: Biochemistry I Fall Term, 2004Documento6 páginasLecture 14: Enzyme Kinetics: Biochemistry I Fall Term, 2004Louis FortunatoAún no hay calificaciones
- Lecture Notes Enzyme 2 Enzyme Kinetics WebDocumento29 páginasLecture Notes Enzyme 2 Enzyme Kinetics WebAldren RebaLdeAún no hay calificaciones
- 10-11, Kinetics of Enzyme Catalyzed ReactionDocumento34 páginas10-11, Kinetics of Enzyme Catalyzed ReactionS. Ansary100% (1)
- EnzymeKinetics by P.C. Misra Professor, Department of Biochemistry Lucknow University, Lucknow-226 007Documento21 páginasEnzymeKinetics by P.C. Misra Professor, Department of Biochemistry Lucknow University, Lucknow-226 007Dr. SHIVA AITHALAún no hay calificaciones
- Michaelis Manten KineticsDocumento8 páginasMichaelis Manten KineticsAnonymous O1xkZINAún no hay calificaciones
- Biology 3601 Biochemistry Enzyme Kinetics Laboratory BackgroundDocumento5 páginasBiology 3601 Biochemistry Enzyme Kinetics Laboratory Backgroundkgeorges27Aún no hay calificaciones
- Michaelis-Menten Type Kinetics: E S) ES) K K K E S) ES)Documento2 páginasMichaelis-Menten Type Kinetics: E S) ES) K K K E S) ES)fintastellaAún no hay calificaciones
- Lecture 12 - Enzyme Kinetics IDocumento30 páginasLecture 12 - Enzyme Kinetics IThomas Jones50% (2)
- Lectures - Biochemistry 1 - 2021-2022 Prof Version 6.01 - 23-24-231-246Documento16 páginasLectures - Biochemistry 1 - 2021-2022 Prof Version 6.01 - 23-24-231-246Ken M'voulaboloAún no hay calificaciones
- Enzyme M&LDocumento6 páginasEnzyme M&LNethra Sasikumar100% (1)
- On The Numerical Computation of Enzyme Kinetic Parameters: Biomath CommunicationsDocumento24 páginasOn The Numerical Computation of Enzyme Kinetic Parameters: Biomath CommunicationsNatalia OchoaAún no hay calificaciones
- Enzyme KineticsDocumento10 páginasEnzyme KineticsQuenneBelocuraAún no hay calificaciones
- Enzyme KineticsDocumento8 páginasEnzyme KineticsronojoysenguptaAún no hay calificaciones
- Dr. Nazir Ahmad: Nehal BilalDocumento15 páginasDr. Nazir Ahmad: Nehal BilalAayat MughalAún no hay calificaciones
- Enzymology (GEB-2101) Lecture 3Documento15 páginasEnzymology (GEB-2101) Lecture 3Niloy GhoshAún no hay calificaciones
- Enzyme and Acid - Base CatalysisDocumento64 páginasEnzyme and Acid - Base Catalysisbinseung skzAún no hay calificaciones
- BiochemDocumento5 páginasBiochemNeil BrionesAún no hay calificaciones
- A8 KineticsDocumento6 páginasA8 KineticsGabby TanakaAún no hay calificaciones
- MIT Lecture 6 5.07 Biochemistry LectureDocumento16 páginasMIT Lecture 6 5.07 Biochemistry LectureakiridoAún no hay calificaciones
- Enzyme Kinetics: Medical Biochemistry, Lecture 24Documento39 páginasEnzyme Kinetics: Medical Biochemistry, Lecture 24jeyankarunanithiAún no hay calificaciones
- Enzyme Kinetics GuideDocumento26 páginasEnzyme Kinetics GuideToga Brandon100% (1)
- Enzymes: Kinetics, Specificity, and RegulationDocumento41 páginasEnzymes: Kinetics, Specificity, and RegulationConorTankGorbatsjovAún no hay calificaciones
- Michaelis Menten DerivedDocumento4 páginasMichaelis Menten DerivedGOOGLI TECAún no hay calificaciones
- Michaelis Menten EquationDocumento9 páginasMichaelis Menten Equationsadaf zaidiAún no hay calificaciones
- CB Chapter3Documento30 páginasCB Chapter3Leidy IracemaAún no hay calificaciones
- Lec 3 Simple Enzyme KineticsDocumento27 páginasLec 3 Simple Enzyme KineticsMohamed AbdelaalAún no hay calificaciones
- Atkins EnzimasDocumento5 páginasAtkins EnzimasConstanza Espinoza LaraAún no hay calificaciones
- MIT5 07SCF13 Lec7 8Documento19 páginasMIT5 07SCF13 Lec7 8Ruben RodriguezAún no hay calificaciones
- Michaelis-Menten Kinetics: Robert Roskoski, Blue Ridge Institute For Medical Research, Horse Shoe, NC, USADocumento9 páginasMichaelis-Menten Kinetics: Robert Roskoski, Blue Ridge Institute For Medical Research, Horse Shoe, NC, USAValeria CazaresAún no hay calificaciones
- Enzyme KineticsDocumento13 páginasEnzyme KineticsKhushbu JainAún no hay calificaciones
- Solutions Chapter8Documento25 páginasSolutions Chapter8Fábio RamalhoAún no hay calificaciones
- AT Unit 3-Part 1Documento53 páginasAT Unit 3-Part 1ShivAún no hay calificaciones
- Article Exact and Approximate Solutions For The Decades-Old Michaelis-Menten Equation: Progress-Curve Analysis Through Integrated Rate EquationsDocumento9 páginasArticle Exact and Approximate Solutions For The Decades-Old Michaelis-Menten Equation: Progress-Curve Analysis Through Integrated Rate EquationsElbahi DjaalabAún no hay calificaciones
- Lecture Notes On BCH 409: Advanced Enzymology (3 Units) : Enzymes and Life ProcessesDocumento21 páginasLecture Notes On BCH 409: Advanced Enzymology (3 Units) : Enzymes and Life ProcessesAkash AroraAún no hay calificaciones
- Michaelis-Menten Kinetics and Briggs-Haldane Kinetics - ChemwikiDocumento15 páginasMichaelis-Menten Kinetics and Briggs-Haldane Kinetics - ChemwikiEmmanuel Alexander PilapilAún no hay calificaciones
- Trump's GOP Is Increasingly Racist and Authoritarian-And Here To StayDocumento10 páginasTrump's GOP Is Increasingly Racist and Authoritarian-And Here To Staycharanmann9165Aún no hay calificaciones
- Be Alarmed - The BulwarkDocumento4 páginasBe Alarmed - The Bulwarkcharanmann9165Aún no hay calificaciones
- They Are What They Say They HateDocumento4 páginasThey Are What They Say They Hatecharanmann9165Aún no hay calificaciones
- Understanding The Trump Voters - Here's Why Nobody Is Doing It RightDocumento7 páginasUnderstanding The Trump Voters - Here's Why Nobody Is Doing It Rightcharanmann9165Aún no hay calificaciones
- 2020 Election - Mike Pompeo's Twitter Account Is Trolling Trump's Behavior - VoxDocumento6 páginas2020 Election - Mike Pompeo's Twitter Account Is Trolling Trump's Behavior - Voxcharanmann9165Aún no hay calificaciones
- Who Are Joe Biden's Top Cabinet Picks - Biden AdministrationDocumento4 páginasWho Are Joe Biden's Top Cabinet Picks - Biden Administrationcharanmann9165Aún no hay calificaciones
- Science Denier Science ProfesorDocumento3 páginasScience Denier Science Profesorcharanmann9165Aún no hay calificaciones
- Former White Supremacist - This Is How To Tackle Hate and Bigotry (Opinion)Documento5 páginasFormer White Supremacist - This Is How To Tackle Hate and Bigotry (Opinion)charanmann9165Aún no hay calificaciones
- The 170 Items That Will Disappear First in A Disaster - Stock Up On These Now!Documento18 páginasThe 170 Items That Will Disappear First in A Disaster - Stock Up On These Now!charanmann9165Aún no hay calificaciones
- World Toilet Day - Checking Wastewater Can Speed Up Covid-19 TestsDocumento3 páginasWorld Toilet Day - Checking Wastewater Can Speed Up Covid-19 Testscharanmann9165Aún no hay calificaciones
- 'They Created A False Image' - How The Reagans Fooled America - Television & RadioDocumento5 páginas'They Created A False Image' - How The Reagans Fooled America - Television & Radiocharanmann9165Aún no hay calificaciones
- Stop Scapegoating Donald Trump. The President Didn't Get Us Into This - by Kitanya Harrison - Nov, 2020 - GENDocumento5 páginasStop Scapegoating Donald Trump. The President Didn't Get Us Into This - by Kitanya Harrison - Nov, 2020 - GENcharanmann9165Aún no hay calificaciones
- 'Deep Down, He's A Terrified Little Boy' - Bob Woodward, John Bolton and Others On TrumpDocumento12 páginas'Deep Down, He's A Terrified Little Boy' - Bob Woodward, John Bolton and Others On Trumpcharanmann9165Aún no hay calificaciones
- Nations Long Targeted by US Chide Trump's Claims of FraudDocumento6 páginasNations Long Targeted by US Chide Trump's Claims of Fraudcharanmann9165Aún no hay calificaciones
- Is It Treason - As Trump Denies Election Results, Legal Scholar Unpacks His Attempt To Stay in OfficeDocumento5 páginasIs It Treason - As Trump Denies Election Results, Legal Scholar Unpacks His Attempt To Stay in Officecharanmann9165Aún no hay calificaciones
- Trump's Labor Secretary Is A Wrecking Ball Aimed at Workers - The New YorkerDocumento20 páginasTrump's Labor Secretary Is A Wrecking Ball Aimed at Workers - The New Yorkercharanmann9165Aún no hay calificaciones
- Stacey Abrams On Minority Rule, Voting Rights, and The 2020 ElectionDocumento21 páginasStacey Abrams On Minority Rule, Voting Rights, and The 2020 Electioncharanmann9165Aún no hay calificaciones
- How Falwell Kept His Grip On Liberty Amid Sexual Games,' Self-DealingDocumento41 páginasHow Falwell Kept His Grip On Liberty Amid Sexual Games,' Self-Dealingcharanmann9165Aún no hay calificaciones
- Expert Details The Secretive Shadow Network' Behind America's Radical Right For The Past 40 Years - Raw StoryDocumento37 páginasExpert Details The Secretive Shadow Network' Behind America's Radical Right For The Past 40 Years - Raw Storycharanmann9165Aún no hay calificaciones
- John Oliver Makes The Case For Impeachment Against Dangerous' Bill Barr If Trump Wins - Raw StoryDocumento10 páginasJohn Oliver Makes The Case For Impeachment Against Dangerous' Bill Barr If Trump Wins - Raw Storycharanmann9165Aún no hay calificaciones
- Aubrey ODay, Donald Trump Jr. Ex, Tweets & Has ReceiptsDocumento11 páginasAubrey ODay, Donald Trump Jr. Ex, Tweets & Has Receiptscharanmann9165Aún no hay calificaciones
- Trump Is The Ultimate Corrupt Washington PoliticianDocumento16 páginasTrump Is The Ultimate Corrupt Washington Politiciancharanmann9165Aún no hay calificaciones
- US Election 2020 - How Trump Has Changed The WorldDocumento18 páginasUS Election 2020 - How Trump Has Changed The Worldcharanmann9165Aún no hay calificaciones
- The Infodemic' of COVID-19 Misinformation, ExplainedDocumento4 páginasThe Infodemic' of COVID-19 Misinformation, Explainedcharanmann9165Aún no hay calificaciones
- The (Almost) Complete History of 'Fake News' - BBCDocumento17 páginasThe (Almost) Complete History of 'Fake News' - BBCcharanmann9165Aún no hay calificaciones
- The Trumpist Death CultDocumento5 páginasThe Trumpist Death Cultcharanmann9165Aún no hay calificaciones
- What We Know So Far About How COVID Affects The Nervous System - Scientific AmericanDocumento9 páginasWhat We Know So Far About How COVID Affects The Nervous System - Scientific Americancharanmann9165Aún no hay calificaciones
- The Trumpist Death CultDocumento5 páginasThe Trumpist Death Cultcharanmann9165Aún no hay calificaciones
- Trump ImplosionDocumento21 páginasTrump Implosioncharanmann9165Aún no hay calificaciones
- Amateur Hour at The Trump White House - POLITICODocumento11 páginasAmateur Hour at The Trump White House - POLITICOcharanmann9165Aún no hay calificaciones
- MICROPAR PPT Group ADocumento43 páginasMICROPAR PPT Group AEben Alameda-PalapuzAún no hay calificaciones
- Approach To Malabsorption (SANJAY)Documento58 páginasApproach To Malabsorption (SANJAY)Sanjay KumarAún no hay calificaciones
- Lession Plan - MIDocumento21 páginasLession Plan - MINithya SannidhiAún no hay calificaciones
- Engineering Mechanics Lectures PDFDocumento83 páginasEngineering Mechanics Lectures PDFluay adnanAún no hay calificaciones
- NNDC Planning Applications 4oct - 11 OctDocumento4 páginasNNDC Planning Applications 4oct - 11 OctRichard SmithAún no hay calificaciones
- Metric Heavy Hex Nuts: ASME B18.2.4.6M-2010Documento16 páginasMetric Heavy Hex Nuts: ASME B18.2.4.6M-2010CarlitosAún no hay calificaciones
- Inkontinensia Urin: Dr. Adhi Permana, SPPDDocumento35 páginasInkontinensia Urin: Dr. Adhi Permana, SPPDTiara KhairinaAún no hay calificaciones
- Proceedings of National Conference on Landslides held in LudhianaDocumento8 páginasProceedings of National Conference on Landslides held in LudhianaAniket PawarAún no hay calificaciones
- Borneo SporenburgDocumento2 páginasBorneo SporenburgDorin TecuceanuAún no hay calificaciones
- Quant One Analyser – endless possibilitiesDocumento6 páginasQuant One Analyser – endless possibilitiesSamuel SuAún no hay calificaciones
- Guide to Conducting SAFOP StudiesDocumento52 páginasGuide to Conducting SAFOP Studiesokemma79% (14)
- Lab 1 Free Fall GEC - CEA21 - OERSTEDDocumento6 páginasLab 1 Free Fall GEC - CEA21 - OERSTEDLee-Ann LimAún no hay calificaciones
- 3 Edition February 2013: Ec2 Guide For Reinforced Concrete Design For Test and Final ExaminationDocumento41 páginas3 Edition February 2013: Ec2 Guide For Reinforced Concrete Design For Test and Final ExaminationDark StingyAún no hay calificaciones
- Technical PaperDocumento6 páginasTechnical PaperSIJO MONACHANAún no hay calificaciones
- Unit explores Christian morality and conscienceDocumento1 páginaUnit explores Christian morality and conscienceRose Angela Mislang Uligan100% (1)
- Classification of Placenta PDFDocumento5 páginasClassification of Placenta PDFAdarsh jainAún no hay calificaciones
- Hydraulic Power Steering System Design PDFDocumento16 páginasHydraulic Power Steering System Design PDFAdrianBirsan100% (1)
- Man FXM FKM Motors PDFDocumento176 páginasMan FXM FKM Motors PDFRenato MeloAún no hay calificaciones
- Vapour Bar Exchange IMFL PackageDocumento4 páginasVapour Bar Exchange IMFL PackageNishank AgarwalAún no hay calificaciones
- Purification of Morphologically and Functionally Intact Human Basophils To Near HomogeneityDocumento9 páginasPurification of Morphologically and Functionally Intact Human Basophils To Near HomogeneitySinaí GutierrezAún no hay calificaciones
- Kirloskar-Oil-Engines DescriptionsDocumento8 páginasKirloskar-Oil-Engines Descriptionssinghhardeep760Aún no hay calificaciones
- Chapter 5 Coordinate GeometryDocumento33 páginasChapter 5 Coordinate GeometryKalAún no hay calificaciones
- Larrabee JChem Educ 1990,67,267Documento3 páginasLarrabee JChem Educ 1990,67,267κ.μ.α «— Brakat»Aún no hay calificaciones
- Civil Engineering Subjects (1st - 5th Year) - 1Documento5 páginasCivil Engineering Subjects (1st - 5th Year) - 1Vincent TayagAún no hay calificaciones
- Mouse Deer and TigerDocumento2 páginasMouse Deer and Tigeralan.nevgan100% (1)
- Chemical reactions and structuresDocumento22 páginasChemical reactions and structuresStormy StudiosAún no hay calificaciones
- Shariff NDocumento4 páginasShariff NKruu ChinnuAún no hay calificaciones
- Craniosacral Therapy For The Treatment of Chronic.10Documento9 páginasCraniosacral Therapy For The Treatment of Chronic.10Marcus Dos SantosAún no hay calificaciones
- Tramadol Drug StudyDocumento1 páginaTramadol Drug Studymilkv82% (11)
- HHG4M - Lifespan Development Textbook Lesson 4Documento88 páginasHHG4M - Lifespan Development Textbook Lesson 4Lubomira SucheckiAún no hay calificaciones